Biological Networks Design Principle: Discovery through Dynamic Optimization
Bambang Adiwijaya, Paul I. Barton, Tomás Lozano-Pérez & Bruce Tidor
What
The emerging field of systems biology includes a strong focus on the network perspective of biological systems. In this work we explore the development and application of tools that can be used to discover structure--function relationships in biological networks, to discover design principles and to design synthetic studies of new and modified networks.
Why
Mathematical optimization provides a powerful set of methods to achieve the goals of network analysis, interrogation, and design, with applications in relating the observed structural differences of biological networks with their functional roles, in understanding trade-offs inherent in biological network design, and in developing therapeutic strategies for intervention and modulation of biological networks.
How
We illustrate these approaches in the context of MAP kinase networks, for which there is a strong foundation of experimental and theoretical study, but about which many questions remain. In particular, subtle and significant differences exist between individual MAP kinase cascades that may correspond to important context-specific functional differences.
To understand MAP kinase networks, we posed the problem as a multiple-objective optimization with embedded dynamics. The dynamics of MAP kinase is modeled as a biochemical network, with dynamic equations expressing mass-action kinetics and other relationships. Optimization was applied to a set of network structures, the optimal networks were examined to elucidate underlying design principles, and multiple objectives were imposed to discover inherent trade-offs. The optimization probes structures and dynamics to understand the effectiveness of networks in producing specific signal-processing characteristics such as signal amplification, response time, and noise filtering.
Progress
We have developed computational methods to optimize the dynamics of biological networks represented by differential algebraic equations, to analyze the optimal results in the presence of local degeneracy, and to extract trade-offs among multiple objectives.
Implementation of the methods to MAP kinase networks reveals multiple strategies that the networks may operate. Furthermore, we discovered the signifcant differences in functiaonl properties of networks with varying circuitry, and significant perfornace advantages in noise reduction and responsiveness that non-linear components of biological networks can produce as compared to reaction networks with linear components.
Future
In the algorithm development, we would like to extend the algorithm to optimize simultaneously the network structure and kinetic parameters of the dynamic models of biological networks. In the application development, we would like to apply the methods to larger sized networks to produce more insights which may be useful for identification of therapeutic targets.
Research Support
This research is supported in part by a Merck/MIT Graduate Fellowship, a Poitras Graduate Fellowship, and the National Institutes of Health.
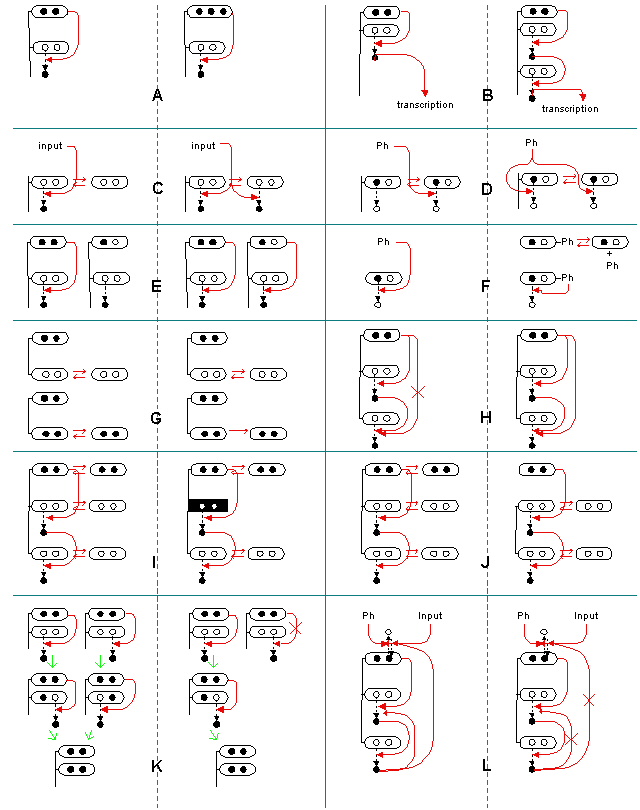
Figure 1: The search space of MAP kinase network structure being probed in this research.
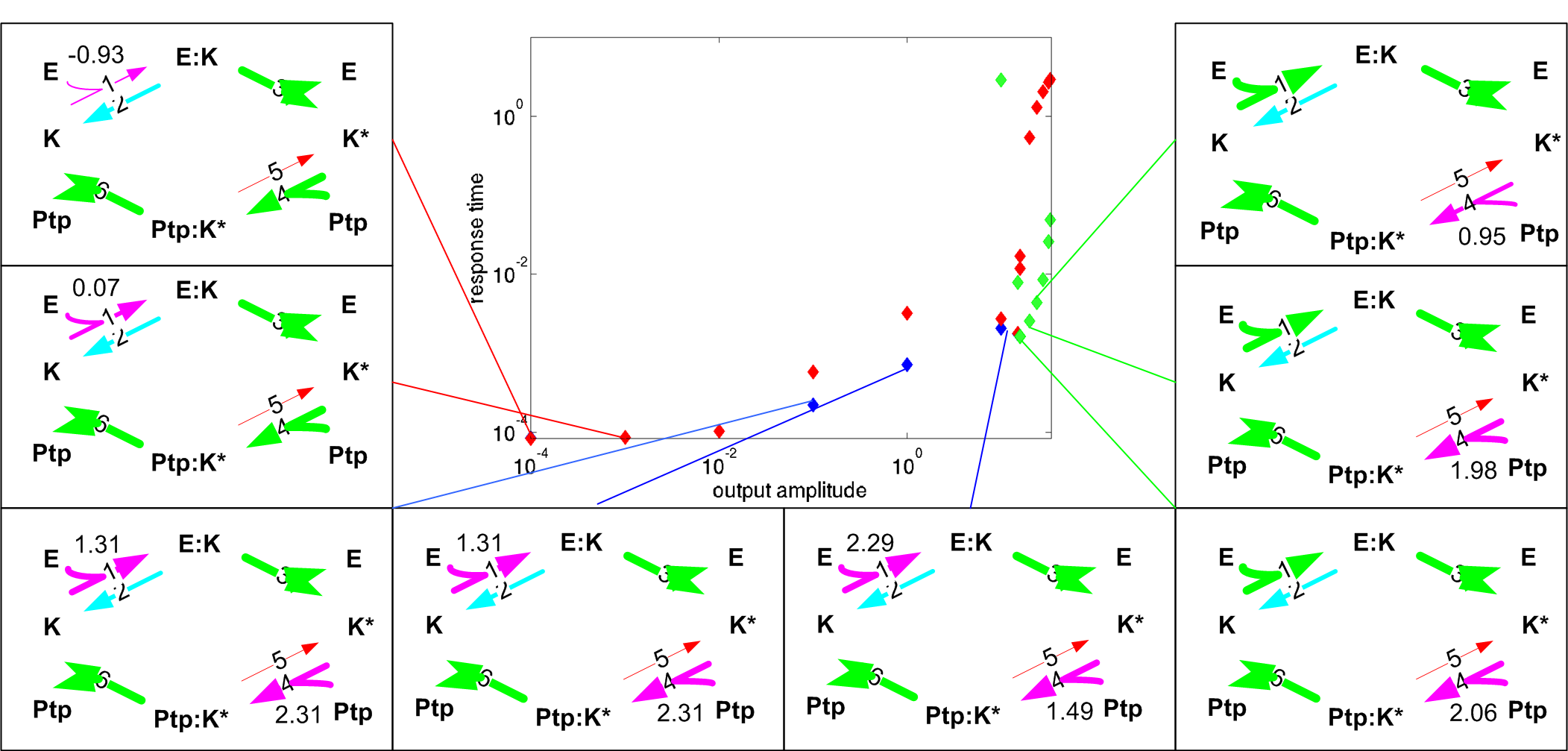
Figure 2: Network optimization Pareto set reveals organization to multiple strategies where the network may operate.
The Stata Center, Building 32 - 32 Vassar Street - Cambridge, MA 02139 - USA tel:+1-617-253-0073 - publications@csail.mit.edu (Note: On July 1, 2003, the AI Lab and LCS merged to form CSAIL.) |
