Re-engineering Enzyme Catalysis Using Computer Modeling and Combinatorial Libraries
Kathryn A. Armstrong & Bruce Tidor
The design and development of protein enzymes that extend the catalytic range beyond the currently available repertoire is important for industrial synthesis. Progress in this area has generally involved making use of structural and/or homologous sequence information of a parent enzyme carrying out a related function that is used to seed a search in sequence space. However, the sequence space available for modification during re-engineering is phenomenally large, and such searches can only examine a tiny fraction of it. Techniques that implicitly explore the larger space to focus on a smaller portion for explicit examination could be particularly useful. The goal of this project is to analyze the evolutionary space computationally and gain insight into the roles of different residues in affecting catalysis, with the ultimate goal of redesigning enzymes more effectively and succeeding in cases where other methods have proved inefficient.
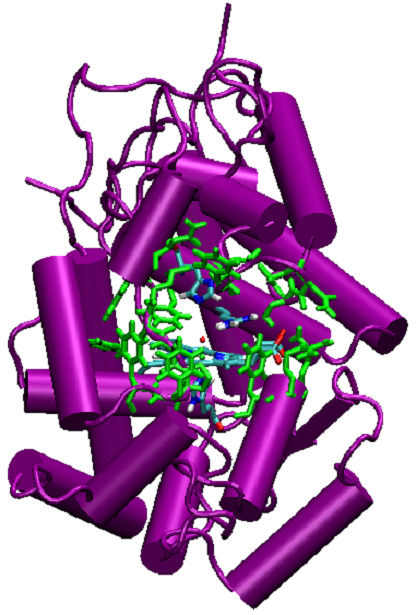
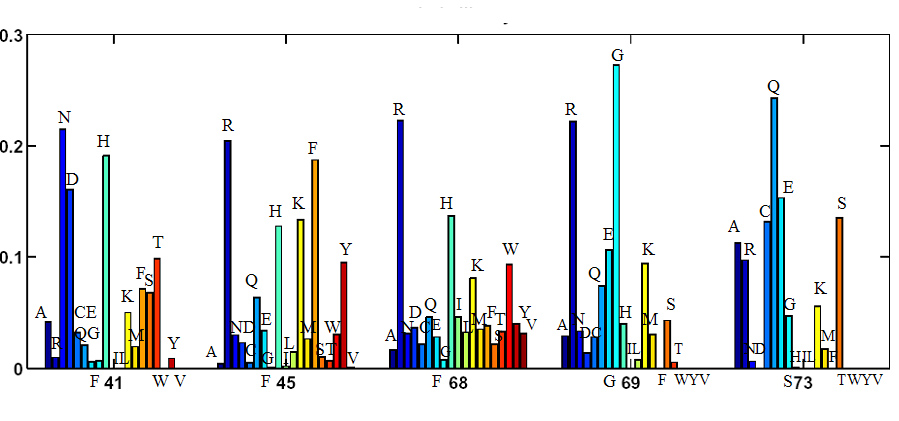
We have computationally analyzed the sequence space of the enzyme horseradish peroxidase both at the active site and at distal locations. This enzyme, shown in Figure 1, is naturally used for the biosynthesis of plant hormones. It is most widely produced for commercial uses, however. Structural information about the enzyme and its mutants is used to examine all sequences at the active site of the enzyme and focus on regions of sequence space that are small enough to be searched with high-throughput experimental techniques. We predict that these regions will be biased toward sequences that are consistent with the native fold, which we hypothesize to be a prerequisite for enzymatic function. Active site residues, highlighted in Figure 1, are analyzed using the dead-end elimination and A* algorithms to produce a rank-ordered list of sequences which are compatible with the backbone fold. The frequency of each amino acid at each of five mutated positions is shown in Figure 2. This distribution is used to create a combinatorial library for testing.
Our mutant libraries merge with experimental efforts underway in the Wittrup and Klibanov laboratories at MIT. The Wittrup lab uses yeast-display and high-throughput screening to completely search the computationally focused regions of sequence space, and the Klibanov laboratory uses advanced enzymology to thoroughly study the kinetic properties of variant enzymes.
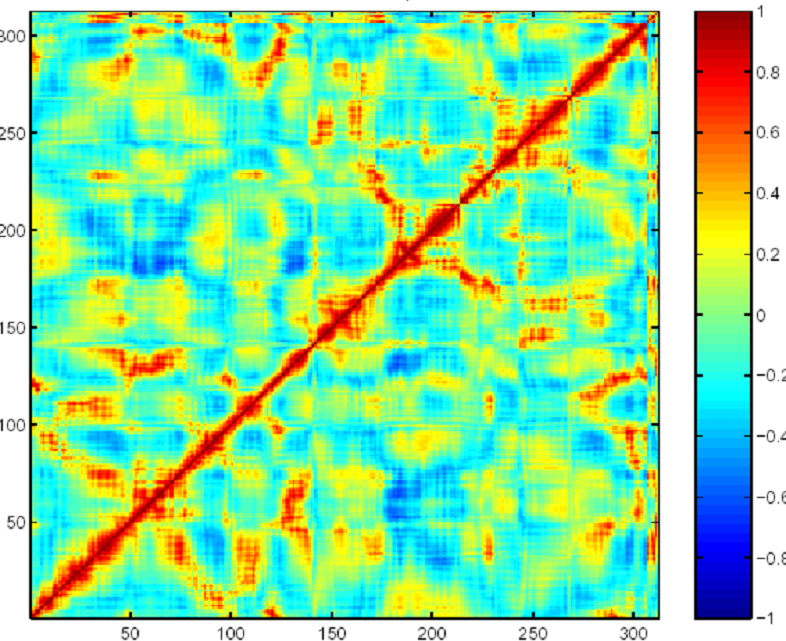
We have also explored the role of distal mutations in the context of dynamics and correlated motions. Revealing distal and active site residues with catalytic influence should offer insight into the evolutionary pressures that created the original enzyme and should give useful suggestions to design effective libraries for testing. Figure 3 shows the results of such analysis. One can see that the active site (roughly between residues 50 and 80) is in fact correlated with residues in other parts of the protein. The residues whose motions are correlated with that of the active site offer useful suggestions toward designing effective libraries for testing.
This joint theoretical and experimental approach will produce understanding of the inner workings of this enzyme through iterative analysis, design, and experimental testing. The techniques developed for this problem may also be generally applicable to other hard protein design problems, involving other functions, in the future.
| Figure 1: Horseradish peroxidase. Active site residues are highlighted in green. |
|||
| Figure 2: Five positions of the horseradish peroxidase were mutated to all possible sequence combinations, resulting in a list of foldable sequences to be screened for catalytic activity. Here we show the frequency with which each amino acid occurs at each position in that list of foldable sequences. |
|||
| Figure 3: Correlation coefficients for the alpha-carbon atoms along the backbone chain of horseradish peroxidase. Red indicates positive correlation and blue indicates negative correlation. |
|||
The Stata Center, Building 32 - 32 Vassar Street - Cambridge, MA 02139 - USA tel:+1-617-253-0073 - publications@csail.mit.edu (Note: On July 1, 2003, the AI Lab and LCS merged to form CSAIL.) |