
| Research Abstracts Home | CSAIL Digital Archive | Research Activities | CSAIL Home |
![]()

|
Research
Abstracts - 2007
|
On The Feasibility Of Approximating Subject Specific Brain Anatomy With An Atlas In Diffuse Optical ImagingAnna Custo, David A. Boas & William M. Wells IIIAbstract:In Diffuse Optical Imaging of brain activity a 3D MRI-based subject anatomical model is used to simulate light propagation in highly scattering tissues. An MRI scan might not always be available or might not be safe to use on particular subjects or in certain situations. In this paper we compare the simulated optical measurements obtained using the subject specific anatomical model versus a general adult human head model. The most relevant measurement, the partial pathlength factor in the brain, indicates that such anatomical approximation can be used without introducing a significant error (relative error <= 10%), provided that a proper atlas is selected. Introduction:We use Monte Carlo (Boas et al. 2002) to simulate the stochastic process of light propagation in an adult human head model of which we specify tissues optical properties such as scattering and absorption coefficients (µa and µs respectively). In brain optical imaging the significant tissues are: skin-type (which includes fat, muscle, connective tissue and scalp), skull-type (which includes bone, outer-table, diploe and inner-table), Cerebral Spinal Fluid (CSF) and brain (which is grey and white matter and glial matter). The adult human head model is generated by segmenting into these four tissue types the subject 3D anatomical MRI scan. Such model creation requires the availability of an MRI scan as well as being able to safely scan the subject, along with a careful analysis of the MRI data to compute the four tissue types segmentation. In this work, we explore the possibility of substituting the MRI-based anatomically correct 3D head model with a general head model, or atlas. We claim that such general anatomical model is sufficient in providing the optical forward model to solve the DOI inverse problem and hence localize brain activation with acceptable spatial accuracy. In order to validate our claim, we simulate measurements of sensitivity to absorption changes in the brain (or partial pathlength factor, PPF) and we calculate the deviation of such measurement against these simulated in the true subject head model. Models:In this work we compare the measurements obtained using a linear probe to simulate photon migration on two anatomical human head models. The linear probe: 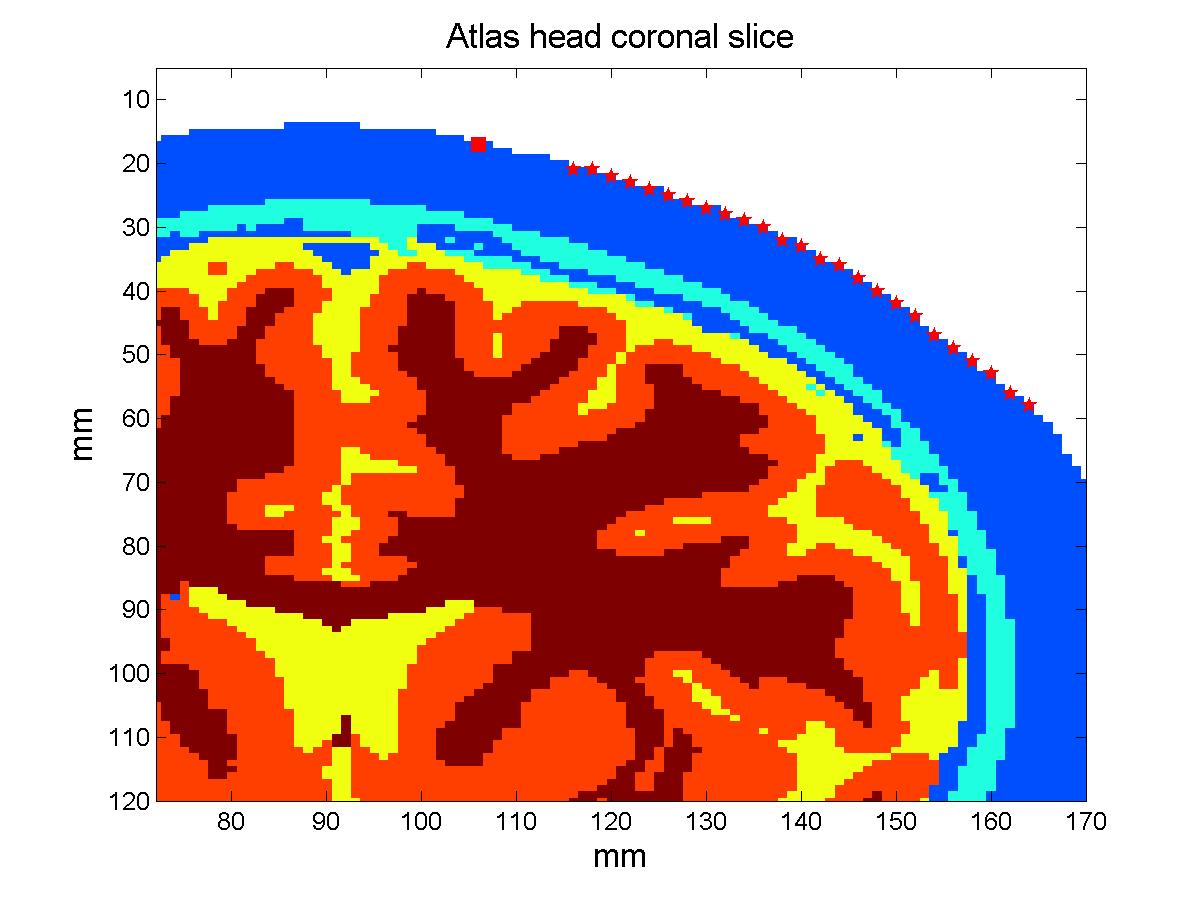
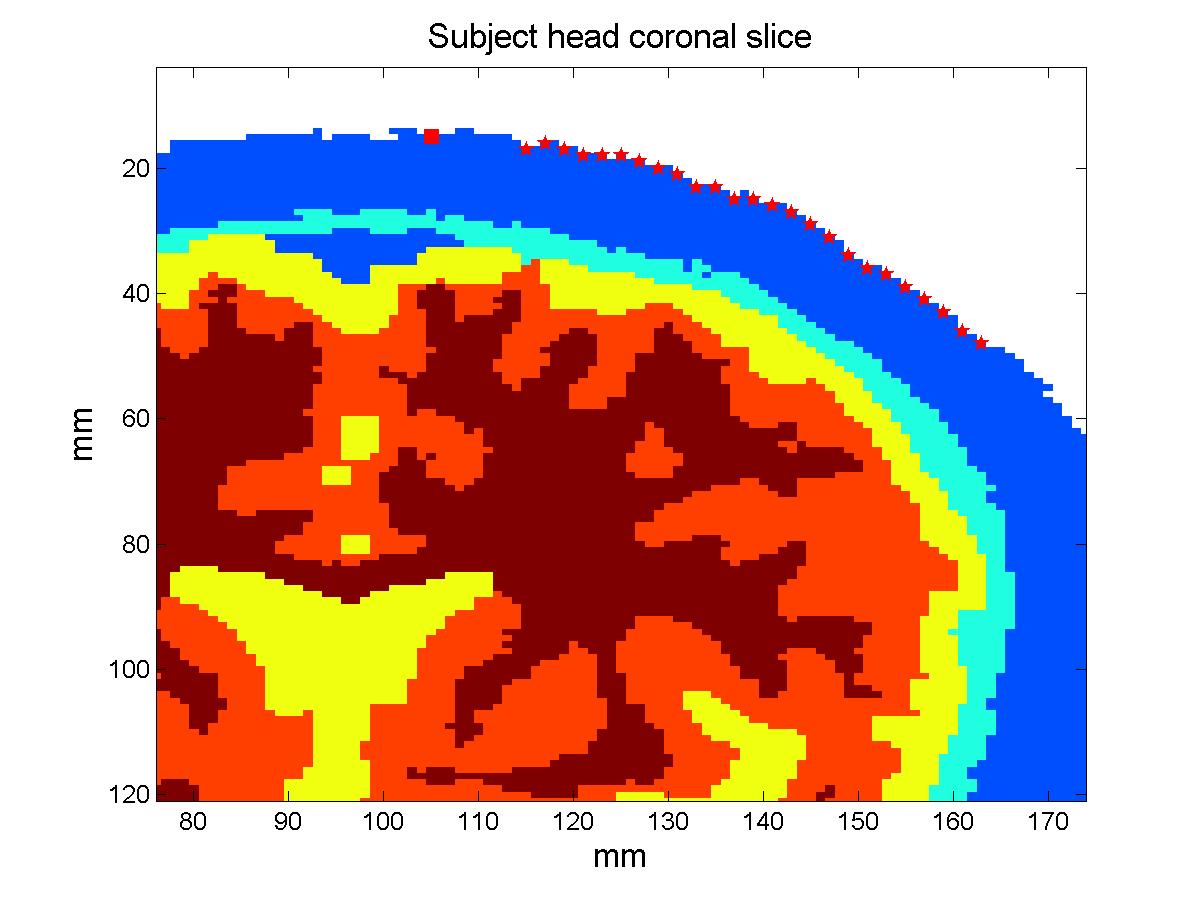
The atlas: The subject model: Methods:In order to compare the measurements that would be obtained simulating photon migration in the subject head with these calculated using the atlas, we need to bring our head structures into the same coordinate system. We can then proceed analyzing the simulated measurements and quantify their deviation. Warping the subject into MNI stereotactic space: The estimated transform is used to find the coordinates of the probe in MNI space and transform the segmented subject head into MNI space, this time employing the nearest neighbor interpolation to resample the voxels labels in the new space. The resulting subject head is a 184x216x184 mm voxel volume. Results:Total and relative PPF: When we look at the relative error plot (Figure 2b), we observe that the sensitivity to µa changes in scalp-skull varies only slightly between the two head models (error less than 4%), in agreement with (Custo et al. HBM 2007), whereas the pathlength factor in the brain varies more, in agreement with the sensitivity to CSF thickness described in (Custo et al. 2006, Custo et al. HBM 2007). However, the PPF relative error is no larger than 10% at separations greater than 25 mm (which is where the photons start to probe the cortex). The error reduces significantly at larger distances due to the longer photon pathlength in the brain and therefore smaller effect of the clear CSF layer (at 36 mm separation PPF relative error approximately equals zero). 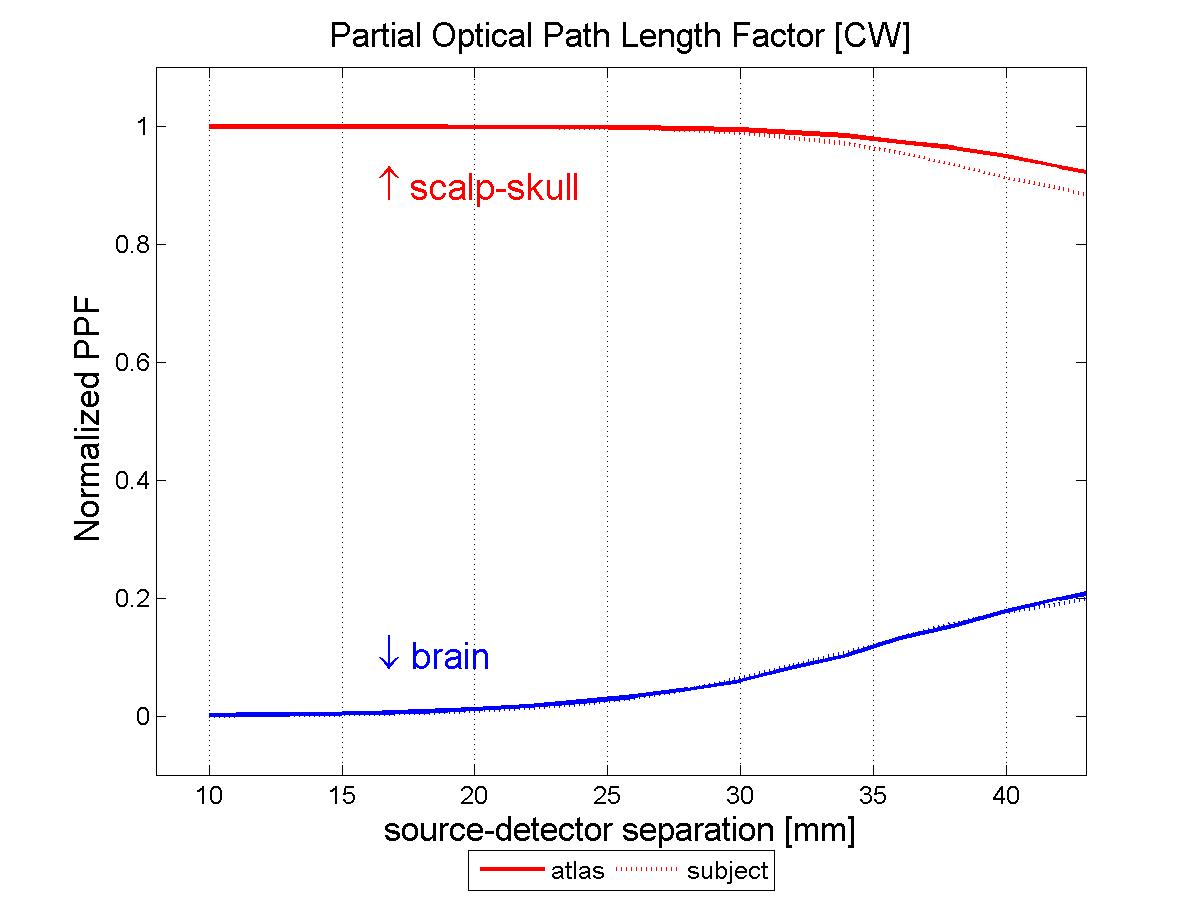
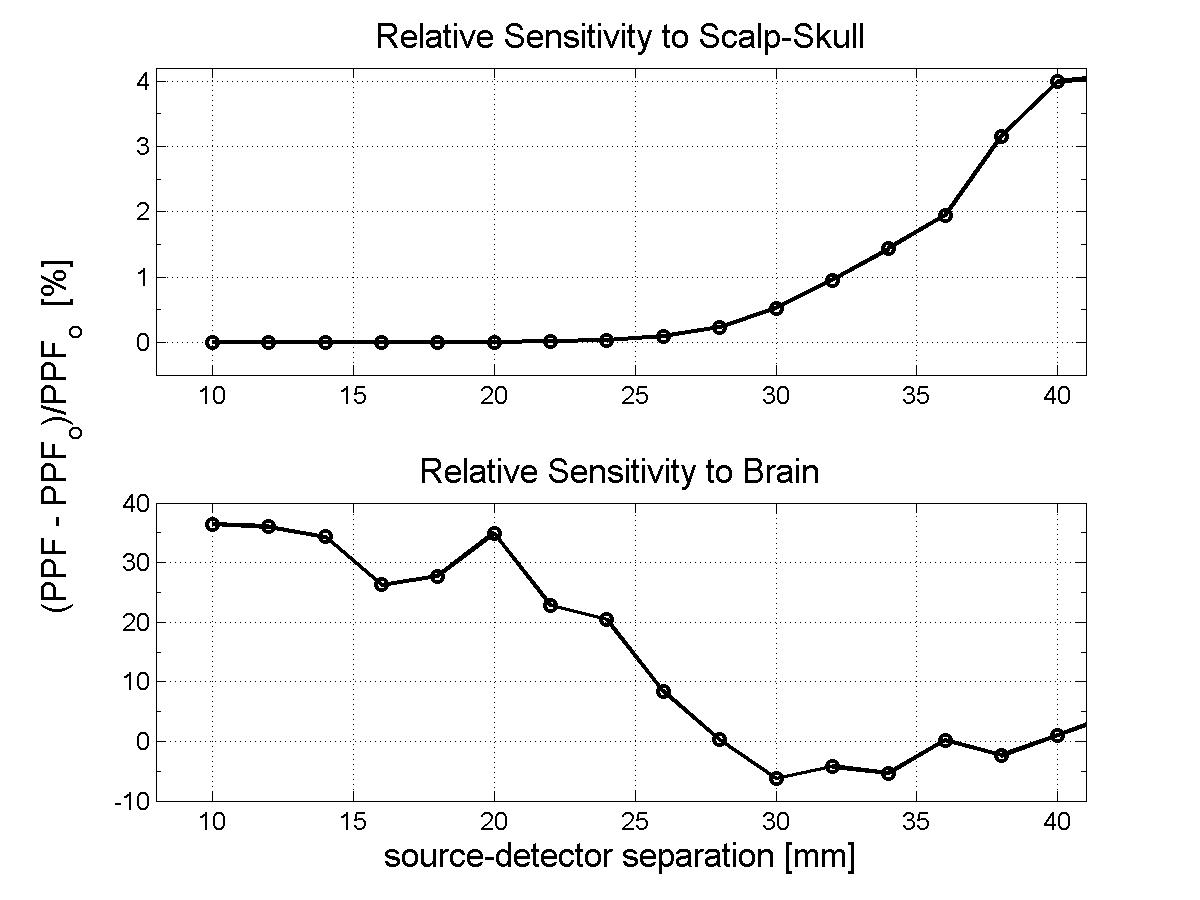
Conclusions:As Figure 2 clearly shows, the results thus far are encouraging. When a proper atlas is selected and the variation of CSF thickness between head models is not significant, the sensitivity to absorption changes in the brain is very well modeled by atlas-based measurements (with error less than 10%). It is left to be establish how best to select a proper atlas. Recent studies have shown variability of cortical normalized volumes and their functional regions across gender. However, so far, there has been little investigation of the variability of CSF across a population as a function of age, gender and race. The general agreement seems to be that the CSF layer increases with age, but we are not aware of any study validating or quantifying such claim. Our next goal is to conduct a study across a large population to characterize CSF variations factors. References:[1] D.A. Boas, J. P. Culver, J. J. Stott and A. K. Dunn. Three dimensional Monte Carlo code for photon migration through complex heterogeneous media including the adult human head. Optics Express vol. 10 pp. 159--170, 2002. [2] D. L. Collins, A. P. Zijdenbos, V. Kollokian, J. G. Sled, N. J. Kabani, C. J. Holmes, and A. C. Evans. Design and Construction of a Realistic Digital Brain Phantom. IEEE transactions on medical imaging vol. 17, no. 3, June 1998. [3] A. Custo, W. M. Wells III, A. H. Barnett, E. M. Hillman, and D. A. Boas. Effective scattering coefficient of the cerebral spinal fluid in adult head models for diffuse optical imaging. Appl Opt vol. 45, no. 19, pp. 4747--55 2006. [4] A. Custo and W. M. Wells III. Effect of CSF thickness in Diffuse Optical Imaging. Human Brain Mapping Chicago, IL, USA, June 2007 |
||||
|